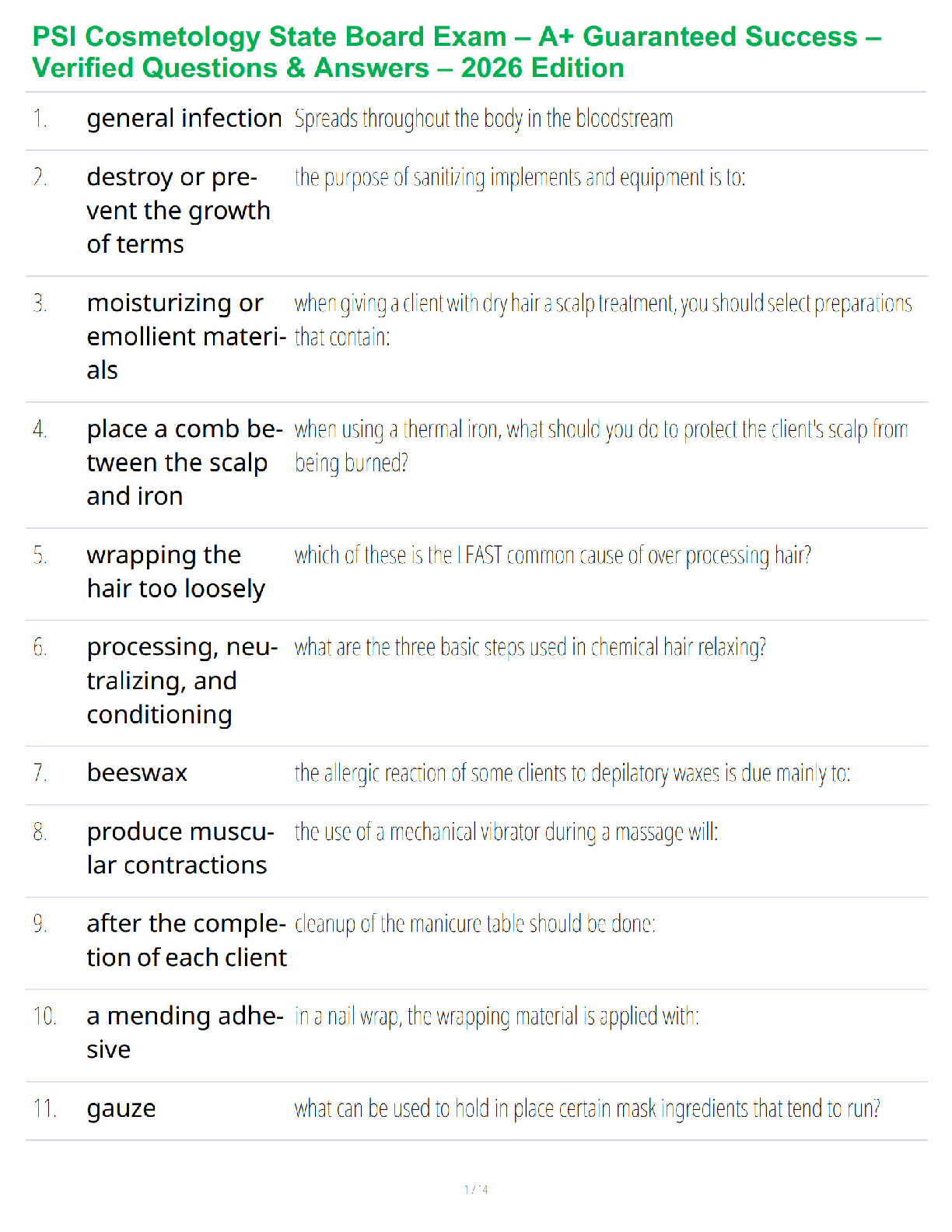
Three energy barriers
Form ES
convert substrate to product
release product
Rate limiting step
step with highest activation barrier
enzyme substrate binding models
lock and key
enzyme complimentary
induced fit hy
...
Three energy barriers
Form ES
convert substrate to product
release product
Rate limiting step
step with highest activation barrier
enzyme substrate binding models
lock and key
enzyme complimentary
induced fit hypothesis
lock and key
favors ES complex
barrier between ES and EP
enzyme complimentary
substrate changes to fit enzyme
induced fit hypothesis
enzyme and substrate change after initial fit
preferred model
types of enzyme catalyzed reactions
general acid base catalysis
covalent catalysis
metal ion catalysis
electrostatic catalysis
catalysis through transition state binding
covalent catalysis
forming covalent intermediate with enzyme
Covalent CatalysisAdvertisement
Side chains of AAs provide nucleophilic centers for attack on electrophilic centers of
substrates.
Form covalent intermediate which can be changed by water or a second substrate for desired
product
Ping-pong kinetic mechanisms
transition state binding
catalysis enhanced by enzyme making optimal conacts to the transition state
irreversible inhibitors
modifies by:
chemical that reacts directly with amino acid or
suicide inhibitor
suicide inhibitor
resembles substrate
goes through steps of catalysis
metal ion catalysis
stabilize s binding
stabilize reaction intermediate redox
interactions (donate electron)
electrostatic catalysis
stabilize s binding
stabilize reaction intermediates
storage polysaccharides
starch
glycogen
starch
found in plants
amylose
amylopectin
d-glucose
Starch
Carbohydrate storage in plantsamylose
long unbranched 1- 4
amylopectin
branched 1-6
glycogen
animal liver
Glycogen
Carbohydrate storage in animal liver
structural polysaccharides
cellulose
chitin
cellulose
plant cells
Cellulose
Structural component of plant cell walls
Glycine
Gly,G
Alanine
Ala,A
Proline
Pro,P
Valine
Val,V
Leucine
Leu,L
Isoleucine
Ile,I
Methionine
Met,M
PhenylalaninePhe,F
Tyrosine
Tyr,Y
pKa: 10.07
Tryptophan
Trp,W
Aspartate
Asp,D
pKa: 3.63
Glutamate
Glu,E
pKa: 4.25
Serine
Ser,S
Threonine
Thr,T
Cysteine
Cys,C
pKa:8.18Selenocysteine
Sec,U
pKa: 5.7
Asparagine
Asn,N
Glutamine
Gln,Q
Pyrrolysine
Pyl,O
Lysine
Lys,K
pKa: 10.53
Arginine
Arg,R pKa:
12.48
Histidine
His,H
pKa: 6.0
Trehalose
Transport/storage in insects
Chitin
Exoskeleton of insects
Hyaluronate
Viscosity, lubrication of extracellular secretions
Proteoglycan
Extracellular matrix of animal tissues
HexokinaseGlucose-6-phosphatase
Phospho-fructokinase
Fructose - 1,6 - bisphosophatase
Pyruvate Kinase
Pyruvate carboxylase
=
Oxaloacetate
=
PEP Carboxykinase
Three steps for dissolving water
1. Break water hydrogen bonds
2. Break solute reactions
3. Make new water-solute interactions
Important interactions involving water (4)
1. Polar compounds
2. Hydration of salts
3. Non-polar substances
4. Amphipathic compounds
How do polar compounds interact with water?
They dissolve in water and form hydrogen bonds with solutes capable
How is water involved in the hydration of salts?
Water interacts with charged solutes (ion-dipole interaction)
When crystalline substances dissolve in water their is an _ in entropy.
increase
How is water involved in reaction with non-polar substances?
Hydrophobic substances such as carbohydrate (C-H weakly polar) tend to aggregate in order to
minimize the amount of surface exposition to water. The water in the vicinity of the solute are
constrained to their possible orientations and form cage-like shells around them.How does water interact with amphipathic compounds?
The compounds form micelles in which their hydrophobic areas are arranged as to expose the
least amount to water. The forces that hold these non-polar regions together are called
hydrophobic interactions.
Why are hydrophobic interactions favored?
Thermodynamic stability, it minimizes the number of water molecules required to
surround the hydrophobic portions of the solute molecules.
Buffers
Aqueous systems that tend to resist changes in pH when small amounts of acid (H+) or base (OH-)
are added. A buffer system consists of a weak acid (proton donor) and its conjugate base (proton
acceptor)
Two examples of water as a reactant
1. Formation/breakdown of ATP (phosphoanhydride bond)
2. Synthesis/breakdown of polypeptides (peptide bond)
Water's important interactions among biomolecules
1. hydrogen bonds
2. ionic interactions
3. van der waals (London-dispersion, dipole-dipole)
4. hydrophobic interactions
Effective Concentration
In polypeptides many interactions occur in close proximity, and are therefore, not totally
independent (AB)Ceff=Kintra/Kinter
Common amino acids
22 different amino acids found in proteins and peptides. Coded for by 3-nucleotide codons in
DNA and RNA and have their own tRNA that is used during peptide synthesis on ribosomes.
Uncommon amino acids (3)
1. modifications
2. different side groups
3. D-amino acids
Uncommon AA due to modificationPost-translational modifications of common residues already part of polypeptide. Typical
modifications: hydroxylations, phosphorylations, and methylations
Uncommon AA due to different side group
Can be intermediates in metabolic pathways or found in small peptides made by bacteria
D-amino acids
Found in small peptides made by bacteria that can have antibiotic properties
To find the pI of a peptide
Use the pKa values directly above and below the pH where the overall charge is neutral
NMR Spectroscopy
purified protein placed in a strong magnetic field and bombarded with radio waves, useful
for proteins not incorporated into crystals
crystal structure:ribbon model of protein
protein is purified and crystallized, then subjected to an intense beam of X-rays. protein model that
shows how the single polypeptide chain folds and coils to form the functional protein, can provide
very detailed atomic information, showing every atom in a protein along with atomic details of
ligands, inhibitors, ions, and other molecules that are incorporated into the crystal
Basis for the ribbon model?
Peptide Bond
Resonance Structures-each peptide bond has some double-bond character and
cannot rotate
a helix is stabilized by _.
hydrogen bonds
What does the ribbon model represent?
The C-N bond in the peptide bond has a partial double bond, prohibiting rotation around it. As a
result, the C,N, and four atoms attached are in the same plane. The peptide chain is therefore
thought of rectangles in the same plane attached by Ca atoms on opposite corners of these
rectangles. It represents the flow of the amino acid chain.
What does the ribbon model represent?The presence of alpha helices and beta sheets
Worm Model
The amino acid backbone is indicated by a tube like structure. Sheets are indicated by flat ribbon
arrows. The helices are indicated by solid tubes or cylinders. This is to emphasize that these
structures are not hollow.
Surface Contour
These structures show what surfaces are available to interact with other proteins and
molecules. The ribbon and worm model might give the impression that their is a lot of empty
space which is not true.
A separate amino acid chain of a protein, several identical or different chains can make up a
protein.
Subunit
Part of a subunit that if separated from the rest of a subunit, still forms a structurally stable unit
Domain
Monomer
Individual subunits
Combination of certain secondary structures like a helices and B sheets that can be found in
several proteins, forming a protein family.
Protein fold
Examples of protein folds
a/B-barrels,a/B sandwiches, B propellers
Dimer
Protein consisting of two monomers/subunits.
Superfamily
large group of proteins/enzymes that all show a very similar folding pattern
Active site
Place in protein structure where substrate can bind.protomer
repeating structural unit in a multimeric protein, can be a single subunit or group of
subunits
How does ammonium sulfate precipitation work?
The solubility of proteins varies according to salt concentration. At low salt concentrations, the
solubility of proteins increases with increased salt concentration. As the salt concentration is
increased further the solubility of the protein decreases. At sufficiently high ionic strength the
protein will almost all be precipitated out from the solution, called salting out.
Describe how salting out works
Salt ions attract water molecules away from protein. As more salt ions are added they bind to
water molecules and the proteins are forced to interact with each other. The proteins aggregate
and precipitate.
Ion-exchange
-separates molecules on the basis of charge, not mass
-beads of the resin are modified so that they contain cationic or anionic functional groups
-a solution that contains the protein of interest is applied to the column containing the resin, and
the sample either binds to resin or passes through the column
-a gradient (e.g., salt or pH) can then be used to elute the desired compound if the compound
adhered to the resin
Size-Exclusion Chromatography
separates proteins on the basis of size
column contains beads with many tiny pores
very small molecules can enter the beads, which slows down their progress, while large
molecules move around/between the beads and thus travel through the column faster
Define hydrophobic chromotography.
the columns are packed with beads that contain hydrophobic side chains which the proteins
interact with
affinity chromatography: specific interactionsbeads in the column have a covalently attached ligand that is designed specifically to attract
specific proteins with an affinity for the ligand
separation based on complimentary biochemical interactions
stationary phase is covalently bound w/ ligands
What does the purification table describe?
How many purification steps are needed. If the protein is pure.
The ratio of enzyme activity relative to total protein is called
.
specific activity
absorbance spectroscopy
measures the concentration of a substance in solution
uses Beer-Lambert law
What does absorbance spectroscopy identify?
Aromatic side chains
SDS-PAGE
SDS is a detergent. It will denature the proteins, and bind to them. Since the SDS molecules are
charged it ends up giving all proteins a reproducible ratio of size to charge. The upshot of all this
is that the position of a protein on the final gel is related solely to its size. Smalley proteins go
faster/farther than large proteins.
Isoelectric focusing
Electrophoresing a mixture of proteins through a pH gradient until each protein stops at the pH
that matches its isoelectric point (pI); because the proteins have no net charge at their isoelectric
points, they can no longer move toward the anode from the cathode.
Four steps of protein sequencing
1. Obtain amino acid composition
2. Determine which amino acid is present at the N-terminus
3. Break disulfide bonds if present
4. React with first fragmenting agent and sequencing of fragments
5. React with different fragmenting agent and sequencing those fragment6. Try to fit all pieces together
7. Repeat (5) with another fragmenting agent if necessary
Trypsin
Cuts the peptide bond at a C-terminal side of the side chain
Asp-N-protease
Cuts the peptide bond at the N-terminal side of the side chain
What can amino acid sequencing tell us?
Can tell us what type of protein the sequenced protein is.
Function of protein
Whether it contains one functional domain or many domains
Within a gene family its shows which amino acids are conserved
Can indicated whether a specific cofactor is bound due to specific order and spacing
What proteins are associated with the membrane through electrostatic interactions and H
bonding with the hydrophilic domainsof integral proteins and the polar head groups
of membrane lipids ?
Membrane proteins
How can peripheral proteins be removed from the membrane?
By altering the pH or salt concentration (ionic strength), removal of a Ca+ group by a chelating
agent, or addition of urea or carbonate
What are firmly associated with the membrane, removed only by agents that interfere with
hydrophobic interactions, such as detergents, organic solvents, or denaturants.
Integral proteins
Covalently attached proteins can be removed from the membrane how?
Phospholipase that releases it from the membrane lipid it is attached to.
Hydropathy Plot
computer analysis of AA sequence
-nonpolar regions --> predited membrane spanning-sequence of >20 AA hydrophobic residues indicative of membrane spanning
Globular Proteins
-Hydrophobic effect dominates formation
-Non-polar inside, polar (charged) outside
-Commonly enzymes and transport proteins
fibrous proteins
-form extended sheets or strands joined by disulfide cross bridges which makes them strong
-made up of repeating a-helices OR B-sheets
-tough and durable
-insoluble structural roles
-common ex. are collagen & a-keratin
membrane proteins
-Transmembrane a-helices and B-sheets
-Tyr and Trp residues found predominantly at water-lipid surface, with other residues in
hydrophobic area
Sequence alignment
Compares two amino acid sequences from homologous proteins to different organisms in order
to identify regions of similarity from a common ancestor
Scoring matrices
Gives an idea about the similarities of AA sequences
Table of all pairwise scores for every combination of AAs
BLOSUM scoring matrices
BLOSUM = Blocks substitution matrices
– Based on BLOCKS database (Henikoff & Henikoff,
1992) of over 2000 conserved amino acid patterns in over 500 proteins
– Based on short conserved sequences (blocks)
How does BLOSUM scoring matrices work?-Replacement of an amino acid penalized with -sign and conservation rewarded with +
sign
-If the mutation does not result in a change of function of protein then the score remains close or
the same
-Replacing a charge (pos-->neg or opp.) can change local structure or function, but if not then
score of 0
What amino acids/sequences are most likely to be conserved?
regulatory sites of enzymes
catalytic sites
amino acids important for structure (cysteine)
hydrophobic/hydrophilic regions
Conserved amino acids
important in the function of enzyme/protein
Evolutionary relationships
Protein sequencing finds a degree of identity between the sequences that can used to make a
distance matrix, which indicates how closely related they are
Based on this a phylogenetic tree can be made
Protein/gene family
Group of evolutionarily related proteins, synonymous with gene family
Common ancestor and typically have similar: 3-D structure, functions, and
significant sequence similarity
Protein Superfamily
Proteins with similar 3-D structure, but generally differ in function and sequence
Proteolytic Cleavage
Modification of the polypeptide chain in which parts of the chain are cleaved off by proteolytic
enzymes and the chain is consequently shortened
Changes at N-terminus
Removal of Met
Acetylation
Addition of fatty acyl groupsChanges at C-terminus
Amidation
Prenylation
Removal of Met
Met is initiating AA in all proteins, which is formulated in prokaryotes
Formyl group is removed
In many residues the Met residue is cleaved off
Acetylation
N-terminus is acetylated
Addition of fatty acyl groups
Fatty acid-associated proteins can be attached to the lipid bilayer of the membrane and are
usually membrane proteins
Can become soluble proteins after fatty acid chain is cleaved
Amidation
Common in peptides/peptide hormones
Enabling full activity
Protects against breakdown
Prenylation
Attachment of isoprenoid:frankly diphosphate or geranylgeranyl diphosphate
Anchor for membrane proteins
Changes in individual AAs
glycosylation
phosphorylation
hydroxylation
sulfation
attachment of lipids
attachment of prosthetic groups
glycosylation
the addition of glucose to blood and tissue proteins; typically impairs protein structure and
function
Phosphorylation added to Ser,The,Tyr,His,Arg,Lys
protein kinase tacks a phosphate group onto the switch protein, in the other direction a protein
phosphatase plucks the phosphate off again.
causes conformational changes, which changes the hydrophobic/hydrophilic profileof the
phosphorylated protein
Hydroxylation
Hydroxyl group added to Pro and Lys
Example collagen
Sulfation
Addition of sulfate group to Tyr
In membrane and secreted proteins
Strengthens protein-protein interaction by the introduction of negatively charged,highly polar
group (more water soluble)
Attachment of lipids to individual AAs
To Cys
G-proteins, seven transmembrane receptors
Prosthetic groups
Cofactors that are permanently bound to the enzyme
FAD,FMN,biotin,metal-contatining cofactors
Denaturation is a loss of _ and _ structure. The _ structure stays intact.
tertiary,secondary,primary
What four things denature proteins?
urea
high temperature
low pH
detergent
How does urea denature a protein?
It disrupts hydrophobic interactions
How does low pH denature a protein?
Causes protonation of side chains Asp,His,and Glu, preventing electrostatic interactionsHow do high temperatures denature a protein?
Provide thermal energy greater than the weak interactions (hydrogen bonds, electrostatic
interactions, hydrophobic interactions, and Van Der Waals) involved, causing them to
break.
How do detergents denature proteins?
The detergents interact with the hydrophobic regions of the protein and therefore the protein
cannot interact with other molecules
Tm
Melting point where [folded]=[unfolded]
Circular dichroism spectroscopy
Measurement of the differences in the absorption of left-handed versus right-handed planepolarized light which are given rise to by structural asymmetry in a molecule. Measures the
amount of helical structure in protein.
How is circular dichroism spectroscopy used?
Equal amounts of left and right-handed circularly polarized light at radiated into a chiral
(circular) solution. One of the 2 types is absorbed more than the other, and this wavelengthdependent difference in absorption is measured, yielding the CD spectrum
DNAK and DNAJ are
chaperone proteins that bind to newly synthesized proteins to slow down folding process
(ATP dependent)
How do chaperons DnaJ and DnaK function in protein folding?
DnaJ binds to regions of unfolded residues on the target polypeptide chain rich in
hydrophobic residues, preventing aggregation.
Polypeptide becomes low-affinity ATP-DnaK complex (DnaJ released)
This stimulates ATPase activity of DnaK,causing a conformational change into highaffinity ADP-DnaK-substrate complex
GrpE binding to DnaK results in disassociation of ADP and binding of ATP
This destabalizes the interaction of DnaK and substrate protein, causing release of
substrate from chaperone
Isomerase function in protein folding
Protein disulfide isomerase- catalyzes the interchange or shuffling of disulfide bonds Peptide prolyl cis-trans isomerase- catalyses the interconversion of the cis and trans isomers
of Pro peptide bonds
Prevent folding of parts of the chain until the whole chain has been synthesized
Chaperones
Provide a protected environment for the protein to fold properly
Chaperonins
Steps of Gro-EL-GroES chaperoning function in order
4,3,1,2,5
Role of ubiquitin in breakdown of cellular proteins
Chain of ubiquitin molecules covalently attached to Lys residue of target protein
3 separate enzymes, E1,E2,and E3 involved in process
A polyubiquintylated protein is first recognized by the 19s regulatory particle andthe
ubiquitin is cleaved off
The protein is then fed through the base complex, is unfolded and digested into short peptides
that are released into the cytosol where they are cleaved into ind. AAs by peptidases
What methods are available to detect the folding/unfolding of a protein/enzyme?
loss of activity
absorption spectroscopy
circular dichroism
proteasome function
◦ degrades unneeded,damaged, or faulty proteins by cutting them into small peptides
◦ reulates protein activity by removing it from the cell when ubiquitin moleculesbecome
attached (by destroying it)
The binding of a ligand is often coupled to a conformational change in the protein that
makes the binding site more complementary to the ligand
Induced FitA model of protein function that pictures the binding site of protein and the ligand
fitting together like a _.
lock-and-key model
Ka=1/Kd what is this relationship?
The larger the Ka (and hence the smaller the Kd),the higher the affinity of the protein for the
ligand
Why do we need proteins to transport oxygen?
oxygen is poorly soluble in aqueous solutions and cannot be carried to tissues in sufficient
quantities if it is simply dissolved in blood serum
diffusion of O2 through tissue is insufficient for long distances
Amino acids are not capable of binding to oxygen, so this capability is provided by _.
hemoglobin
Why does oxygen bind to heme C and not a single Fe ion?
The electron donating character of the N-ligands prevent the Fe2+ from being oxidized when
O2 binds, the binding of oxygen to a single Fe2+ ion would result in the formation of reactive
oxygen species that can damage biological structures.
A _ residue is able to form a hydrogen bond with the bound oxygen, increasing the affinity for
oxygen relative to that of free heme C,
histidine
In a multi-subunit protein, a conformational change in one subunit often affects the
conformation of other subunits
Allosteric Effect
What is the change in hemoglobin affinity for molecular oxygen?
Bohr Effect
The hemoglobin tetramer shifts between the _ affinity T-state and high affinity _-state.
What is this shift due to?
low,R,result of a structural change in the tetramer structureHow do different pHs affect hemoglobin?
Low pH- several AAs get protonated and get a positive charge and can now form salt bridges
with other negative AAs. These bridge are only formed in the T-state and trap the enzyme in
this state. The R/T equilibrium is shifted towards the T-state.
High pH-opposite
Why are enzymes such good catalysts?
Accelerates chemical reaction rates
Function in aqueous solutions under very mild conditions of temperature and pH
Highly specific for its substrate and the reaction it will catalyze
To provide for and balance the needs of the cell, enzyme activity is regulated
Transition State vs. Intermediate
Transition states have partially formed bonds and have partially charged atoms.
Intermediates have fully formed bonds.
Michaelis-Menten Equation (equation)
V0= (Vmax * [S]) / ([Km+ [S] )
V0= rate of reaction at a given substrate concentration
Vmax = maximum reaction rate
[S] = substrate concentration
Km = Michaelis constant, represents the [S] at which 1/2 Vmax is achieved
What are the benefits of measuring the initial rate of a reaction V0?
Changes in [S] are negligible, so [S] can be treated as a constant
What does the steady state assumption, as applied to enzyme kinetics, imply?
It assumes the formation of an enzyme-substrate complex that is formed and broken down at
equivalent rates
Catalytic Efficiency
-If [S] << KM, then V0 = kcat/KM[E]T[S]-The rate law resembles a second-order reaction, with kcat/KM being the second-order rate
constant
-In this situation, the substrate binding enzyme becomes the rate-limiting step;
kcat/KMcannot be faster than the frequency which the enzyme and substrate collide
competitive inhibitor
competes with a substrate for the enzyme-substrate binding site
inhibitor can be overcome with high enough substrate concentrations
Inhibitors that bind covalently with or destroy a functional group that is essential for the
enzymes activity
Irreversible Inhibitor
Reversible Inhibitor
competitive and non-competivie inhibitors
Mixed Noncompetitive Inhibitor
Binds at a site distinct from substrate active site, but it binds to either E or ES
Influences the binding of S at active site
Acid-Base Catalysis
Catalysis in which a proton is transferred in the transition state
Catalysis involves H+ or OH- diffusion into catalytic center
Specific acid-base catalysis
Involves acids and bases other than H+ and OH-. These other acids and bases facilitate
transfer of H+ in transition state.
general acid-base catalysis
Catalysis Mechanisms: Metal Ion Catalysis
Multiple possible functions:
1. Serves as electrophilic catalyst by stabilizing -charge intermediate or increased
electron density
2. Generate nucleophile by increasing the acidity of a nearby molecule (i.e. H20)
3. Bind to the substrate, increasing the number of interactions with the enzyme andthus the
binding energy
noncompetitive inhibitorinhibitor binds to another site of enzyme, either E or ES when there is no substrate
uncompetitive inhibitor
binds to enzyme ONLY after substrate binds, only binds to ES complex
Sequential/Single Displacement Reaction leads to the formation of a _.
ternary complex
Double displacement reactions proceed via the formation of _.
a covalently modified enzyme intermediate
What is the turn over number?
number of substrate molecules converted to products per sec by a single enzyme under
optimal conditions and when enzyme is saturated with substrate
Calculated as the initial velocity of the catalyzed reaction (Kcat) at [S]>>Kmdivided by the
enzyme concentration
Three steps of glycolytic pathway that differ from gluconeogenesis
Conversion of glucose into glucose-6-phosphate
Conversion of fructose 6-phosphate into fructose 1,6-bisphosphate
Conversion of phosphoenolpyruvate into pyruvate
Why is gluconeogenesis catalyzed by different enzymes?
Glycolysis is exergonic with ΔG°'=-74 kJ/mol mainly due to the 3 steps that differ, the other
steps are reversible under cellular conditions
For gluconeogenesis to be thermodynamically favorable enzymes must catalyze
reactions
Steps of glycolysis/gluconeogenesis that differ
In glycolysis an ATP used in step 1 and 2, to make the reverse process favorable the reaction
is uncoupled from ATP synthesis and a single Pi is released instead.
In step 3 in glycolysis ATP is produced, to make the reverse step possible an ATP and GTP are
hydrolyzed to put in additional energy to make the step thermodynamically favorable in the
opposite direction.
Main reactive oxygen species
superoxide, hydrogen peroxide, and hydroxyl radicalsWhat role does the pentose phosphate pathway play in removal of reactive oxygen species?
Production of NADPH in the glucose-6-phosphate dehydrogenase reaction regenerates GSH
from its oxidized form GSSG. Reduced GSH protects the cell by destroying free hydroxyl
radicals. Also the co-substrate for glutathione peroxidase that removes hydrogen peroxide. In
both cases, GSH is oxidized to GSSG.
Structural role of sugar
Sugars are the framework of DNA & RNA: The basic building blocks contain a phosphate
group, a ribose (or deoxyribose), and a base. These form the repeating units in RNA and DNA
chains. Linear chains of glucose or derivates form the structural elements in the cell walls of
bacteria and plants.
What about sugars makes it them good structural features of fibers?
The B linkages that form straight chains between sugar molecules is optimal for fibers with high
tensile strength.
Bacterial Cell Wall
Additional peptides are added to further crosslink the carbohydrate chains
Sugar Code
Select proteins and all tissues and cell types have unique carbohydrate units that can be used in
their identification
Last 2 steps of enzyme hydrolase
Residue is recognized by special receptor (or lectin)
When a section of the Golgi complex containing this receptor buds off to form a transport
vesicle, proteins containing manner phosphate residues are dragged intothe forming bud by
integration of the receptor and man. phos.; the vesicle then moves to and fuses with a
lysosome, depositing its contents within
1st 3 steps of enzyme hydrolase
Hydrolase protein contains a signal sequence recognized by SRP particle and as a result the
enzyme is guided to ER during synthesis
In ER lysozyme is glycosylated
Lysosomal enzymes contain a single patch that is recognized by an enzyme that
phosphorylates a mans residue at the terminus end of an oligosaccharide chainHydrolysis of ATP: Charge separation
Charge separation due to hydrolysis relieves electrostatic repulsion among the four negative
charges on ATP. The phosphorous atoms are electron withdrawing groups (partial positive) and
destabilize the molecule with respect to hydrolysis products
Hydrolysis of ATP:Resonance
The product inorganic phosphate (Pi) is stabilized by formation of a resonance hybrid, in which
each of the 4 phosphorous-oxygen bonds has the same degree of double-bond character and the
hydrogen is not permanently associated with any one oxygen.
Hydrolysis of ATP:Ionization
Product ADP2-immediately ionizes, releasing a proton into a medium of very low [H+] (pH7).
The entropy of the solution increases because the more particles, the more disordered the system.
Hydrolysis of ATP:Standard Conditions
The ΔG' value is highly dependent on ATP,ADP,Pi,H+ (pH) and Mg2+. Deviation from the
standard condition causes the value of ΔG' to be more negative.
Hydrolysis of ATP: Solvation
A greater degree of solvation (hydration) of the products Pi and ADP relative to ATP, which
further stabilizes the products relative to the reactants
What does the pentose phosphate pathway create?
Ribose and NADPH
How does the pentose phosphate only create NADPH or ribose?
When both r5p and NADPH are needed by the cell, the first 4 reactions of the pathway
(oxidative steps) predominate. NADPH is produced and r5p is the principal product of carbon
metabolism.
When more r5p is needed than NADPH, the oxidative reactions of the pathway are
bypassed, Withdraw of fructose-6-phosphate and glyceraldehyde-3-phosphate (not glucose-
6-phosphate) from glycolysis and conversion into r5p via a reversal of the transketolase
and transaldolase reactions
How does bifunctional enzyme phosphofructokinase-2/fructose- 2,6-bisphosphatase and product
fructose-2,6-bisphosphate regulate glycolysis and gluconeogenesis Fructose-6-phosphate activates PFK-2 and inhibits F-2,6-BPase
Production of F-2,6-BP stimulates glycolysis by allosteric activation of PFK-1 and
inhibits gluconeogenesis by allosteric inhibition of F-1,6-BPase
Phosphorylation by cAMP-dependent protein kinase inhibits PFK-2 activity and
stimulates F-2,6-BPase
What happens if a muscle preparation containing glycogen phosphorylase is treated with:
phosphorylase kinase and ATP
Conversion of glycogen phosphorylase to the more active, phosphorylated form, phosphorylase a,
glycogen breakdown accelerates.
What happens if a muscle preparation containing glycogen phosphorylase is treated with:
PP1
Converts the active phosphorylase a to the less active phosphorylase b; glycogen break down
slows
What happens if a muscle preparation containing glycogen phosphorylase is treated with:
epinephrine
epinephrine causes the synthesis of cyclic AMP, which activates phosphorylase kinase. The kinase
converts phosphorylase b to a; glycogen breakdown accelerates
How is glycolysis regulated by levels of ATP/AMP
When the cell has ample ATP (low AMP) glycolysis will be inhibited
How is gluconeogenesis regulated by levels of ATP/AMP?
Ample ATP and it is stimulated
What activates or inhibits phosphofructokinase?
activates: AMP (reverse inhibition of ATP), F-2,6-BP
inhibit: ATP, Citrate, decreased pH (lactic acid buildup)
Glycogen synthesis
Activation of UDP-glucose, a reaction catalyzed by UDP-glucose
phosphorylase. Glycogen synthase uses UDP-glucose as a substrate and add the glucose units
to growing glycogen chain. Branches introduced by a branching enzyme. It transfers 6 or 7
residue segments of a growing glycogen chain to the C-6 hydroxyl group of a glucose residues
on the same or nearby chainGlycogen breakdown
Enzyme glycogen phosphorylase cleaves glucose from the nonreducing ends of glycogen
molecules and forms glucose-1-phosphate. It can only do so to long chains and limit dextran are
broken down by a debranching enzyme. In the first step a trisaccharide group from a limit
dextran branch is transferred to the end of a nearby branch. The remaining glucose unit from the
branch is cut off. The glucose-1-phosphate is converted into glucose-6-phosphate by
phosphoglucomutase.
anabolic pathway
•Anabolic pathways consume energy to build complex molecules from simpler ones
•The synthesis of protein from amino acids is an example of anabolism
catabolic pathway
pathway that releases energy by breaking down complex molecules to simples
compounds (polymers to monomers)
ex: cellular respiration
How is NADH recycled under anaerobic conditions?
NADH reduces pyruvate to lactate and is thereby recycled to NAD+
How is NADh recycled in aerobic conditions?
NADH passes electrons to O2
[Show More]










.png)